GC IMAGE - GC Image GCxGC Edition - User Guide 2025R3
New Features and Improvements in Version 2.7
GC Image GCxGC Edition 2.7 has many new features and
improvements. See the GC Image GCxGC Edition
User Guide for full documentation.
What's New in Version 2.7 Release 3 (February 2018)
Improved Support for AB SCIEX .Wiff Files
GC Image GCxGC Edition 2.7 Release 3 includes the following improvements for importing AB SCIEX .Wiff data files:
- New centroiding option for importing Wiff MS1 profile spectra.
- Better performance for importing a Wiff file with multiple samples, and importing and merging multiple Wiff files.
- Random socket exceptions during importing Wiff files are completely eliminated.
In addition, the software includes the following improvements for Import and Merge:
- A progress dialog is now displayed when opening many files to be merged.
- The recently used Cut-by-Cut time settings are now remembered.
New Improvements for Peak Detection, Identification, and Review
GC Image GCxGC Edition 2.7 Release 3 has improved and new features for peak detection, compound identification, and review.
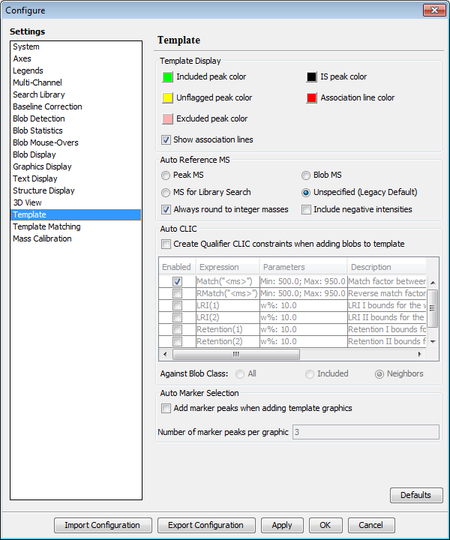
The software includes new settings to control how a reference MS is extracted and used when adding a bob to the template:
- New options to specify the type of spectra to be extracted, Peak
MS, Blob MS, or based on the library search settings. A new
selection tag @lib is introduced to enable CLIC
functions to use the spectra based on the library search settings
during template matching if selected.
- New option to specify if m/z values will be rounded to
integer when extracted. Turning off the rounding option allows
extracting and storing high-resolution m/z values in a reference MS.
- New option to specify if negative intensities will be
included in a reference MS.
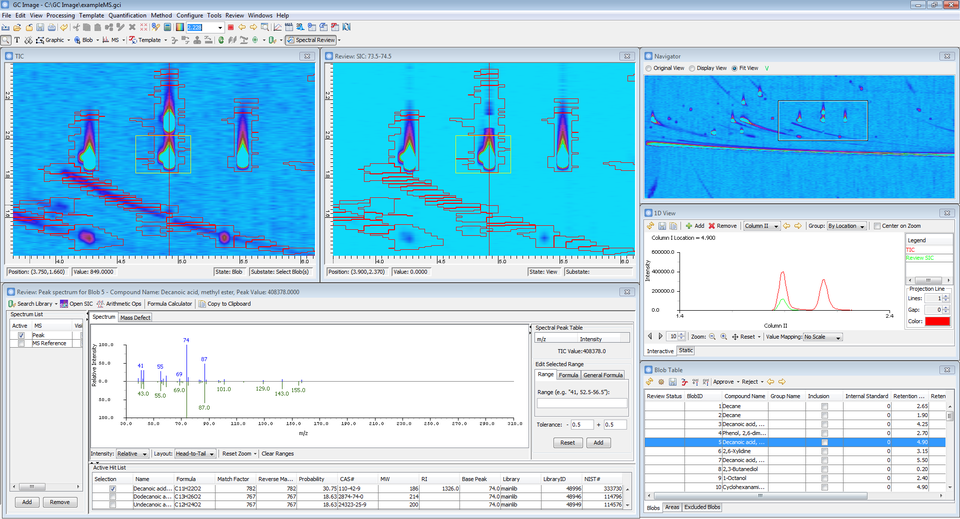
In addition, the software includes new improvements to assist spectral review:
- Reference MS and Hit List are shown in Review Mode's MS Viewer if available.
- The software now position 1D Projection to the Quantifier or Qualifier peak location based on the Review SIC setting.
- The software now detects 0 Min in the user-specified value mapping and ignores negative regions when automatically adjusting color map for SIC.
New Improvements for Multi-Sample Analysis with Investigator
GC Image GCxGC Edition 2.7 Release 3 includes the following improvements on Investigator for multi-sample analysis:
- PCA results are now kept when switching between analysis attributes.
- The Scores and Loadings tables are now saved with the PCA chart report.
- A new option for exporting the filtered feature template based on user selection.
- String values of a CLIC-based attribute now can be saved to an analysis data folder (.GCA).
In addition, the software has a new CLIC function MSString() that allows users to export blob spectra as part of Blob Table
to Investigator or external programs.
What's New in Version 2.7 Release 2 (October 2017)
Improved Support for Heart-Cutting and Variable Modulation
GC Image GCxGC Edition 2.7 Release 2 has improved support for small
chromatograms, such as ones acquired with heart-cutting including:
- Support zooming into tiny regions on 2D View.
- Allow arbitrary scales to be set through the tool bar.
- Allow more projection lines to be used by 1D View.
- Allow creating an edge-to-edge graphical area (rectangle,
polygon, and polyline) for manual integration on chromatograms of any size.
For chromatograms acquired with variable modulation, the software has the following improvements:
- Image Attributes now displays a Variable Modulation table if available.
- Column I dimensions of template graphics are now preserved after saving.
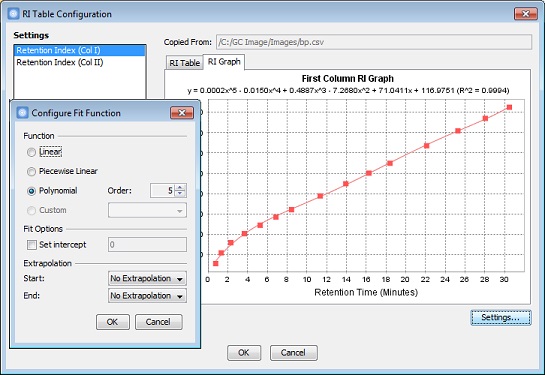
Improved Support for Retention Index Calibration
GC Image GCxGC Edition 2.7 Release 2 includes improvements to support
generic retention index calibration.
- RI tables can be populated from areas and selected blobs in addition to included blobs and a blob set.
- An external RI table can be imported and appended to the existing table.
- The calibration function and R-squared value are shown in the title of RI Graph.
- RI tables have better error checking before applying changes.
If an error is found, the problem index row is automatically selected.
New Improvements for Mass Spectral Viewing and Calibration
GC Image GCxGC Edition 2.7 Release 2 has new improvements for
streamlining the work flow for mass calibration
including:
- New option to open the chromatogram saved by
Export Recalibrated Centroid Image or Export Profile Data to Centroid.
- New OK and Export... button in Configure > Mass Calibration Model that
allows quick export of a recalibrated chromatogram.
In addition, the software has new improvements for the default MS mode including:
- The default MS mode can be set to either
Context, Point, Peak, or Blob/Area through Configure > Settings > Multi-Channel.
- MS Context mode now shows the Point spectrum in addition to Peak and Blob/Area spectra.
- MS Context mode now shows recalibrated spectra for all spectra.
- Improved default color scheme for spectra in the MS Viewer that make different types of spectra more distinct.
New Library Search Features
GC Image GCxGC Edition 2.7 Release 2 includes significant improvements for Library Search features.
- Support In-source HiRes (Identity) search type for Batch search mode.
- Support NIST Library Search Presearch settings: Default, Fast, and Off
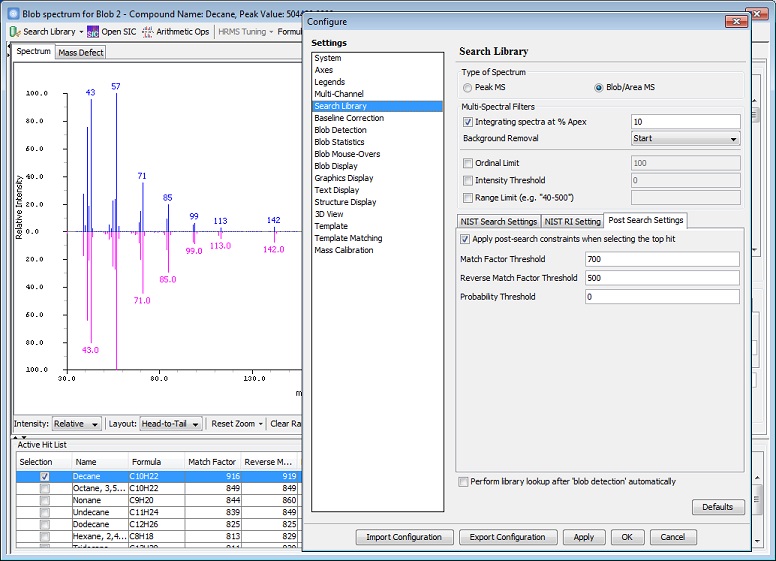
- New Multi-Spectral Filter settings for Search Library:
- Integrating spectra at % Apex: Integrate spectra with TIC values reaching at the specified percent of the apex of a blob or an area.
- Background Removal: Options for None, Start, and Start and End: Determines which portion of the blob to be used as a baseline for correcting library spectra.
- Ordinal Limit: Limit the search spectra to the largest number of intensities.
- Intensity Threshold: Limit the search spectra to intensities greater than the specified threshold.
- Range Limit: Limit the search spectra to the specified spectral range.
- Blob's current Hit List is loaded in MS Viewer when viewing a peak or blob spectrum.
- New Select column in the MS Viewer's Hit List table that clearly shows which hit is selected.
- Non-existent RI values in library search results are now displayed as empty instead of -1.
 |
What's New in Version 2.7 Release 1 (June 2017)
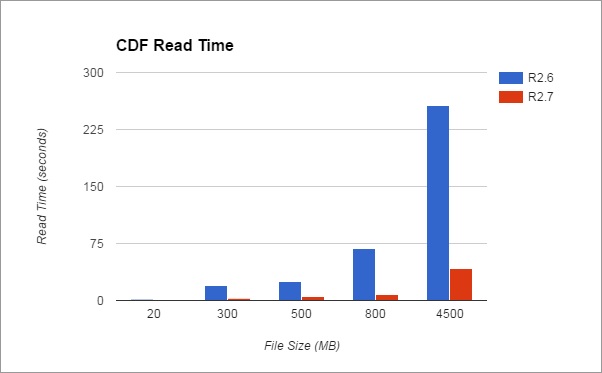
Large Data File Support
GC Image GCxGC Edition 2.7 Release 1 includes significant improvements for reading and writing
large CDF files, which are used as the file store for MS data.
- The software now supports reading and writing of
larger CDF files that can contain more data than previously allowed.
The maximum data size is now only limited by the file
system and memory of the computer.
- Additional improvements yield faster read times for
CDF files on hard disk drives. GC Image now can open
large CDF files much faster!
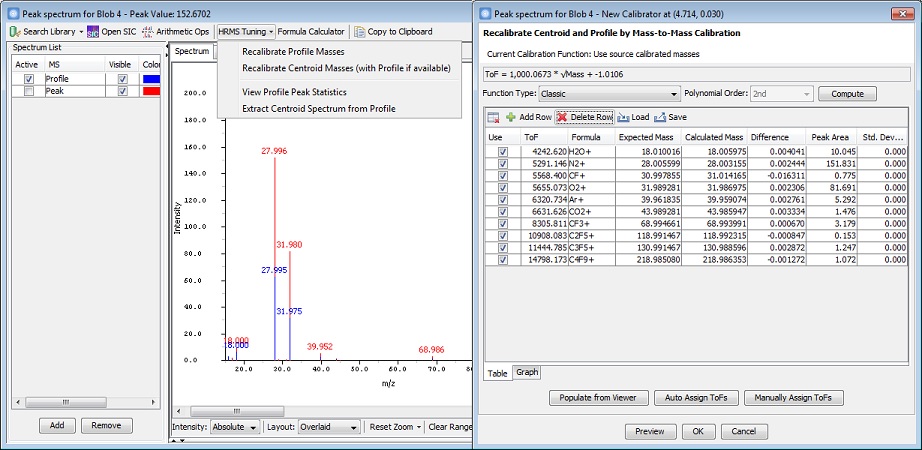
Mass Spectra Centroiding, Calibration, and Viewing
GC Image GCxGC Edition 2.7 Release 1 has new and improved operations for
mass calibration with HRMS data, including:
- The software supports exporting centroid data
extracted from profile data directly to a GCI file (with accompanying CDF
file), in addition to the option of exporting to a standalone CDF file.
- The software supports recalibrating centroid data
with or without profile data, in addition to recalibrating the profile data only.
After recalibration is configured, users can export
the recalibrated centroid data.
- The software has a new, default, context-based MS mode
that opens MS View for point, blob, or area spectra based on content.
If the mouse-click location is not within a blob or area, a point
spectrum is opened; otherwise, both the apex and summed spectra
for the blob or area are opened. If the location is within more
than one object, a popup menu allows selection.
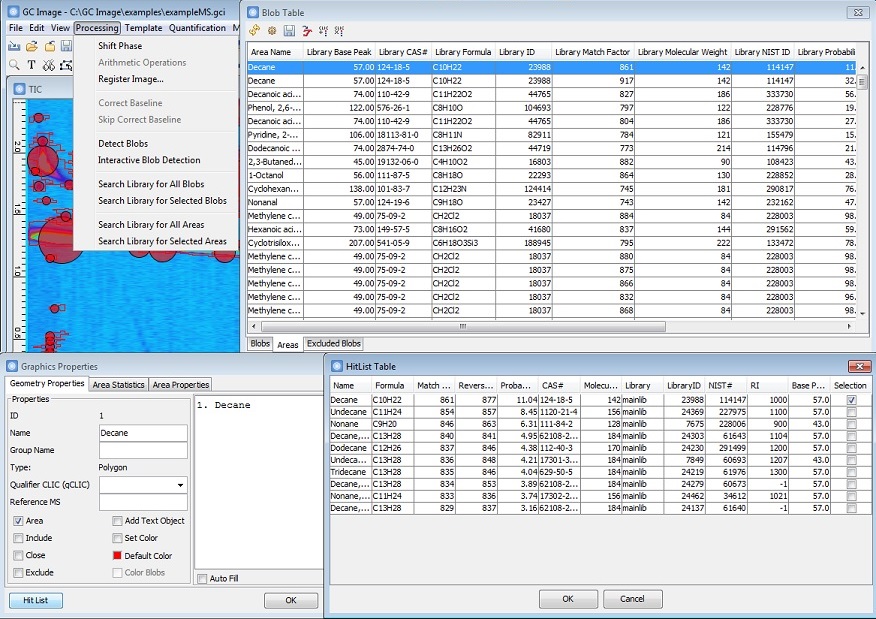
MS Library Search for Area Objects
GC Image GCxGC Edition 2.7 Release 1 introduces MS library search supports for areas.
- New Library Search actions directly search
for matches with spectra from all or selected areas.
- MS Viewer now can apply the hit list from a manual library search to
an area.
- Area properties now include a Hit List that allows
users to examine search results and select the preferred match.
- Library properties now can be added to the Areas
tab of the Blob Table.
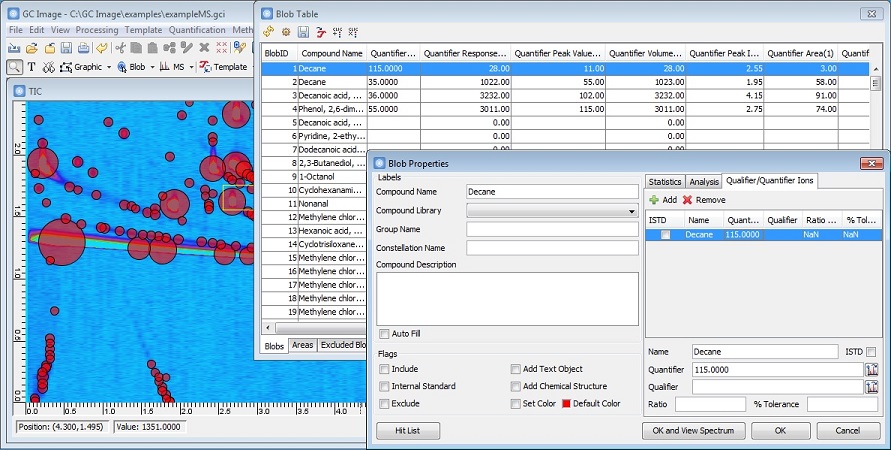
New Quantifier Ion Properties
GC Image GCxGC Edition 2.7 Release 1 has new properties and improved
operations for quantifier ions, including:
- A quantifier ion can be set as an internal standard for blobs and
areas for calibration and quantification, to support
isotope dilution analysis.
- New blob/area statistics for quantifier ions include retention
times, area, peak value, and volume.
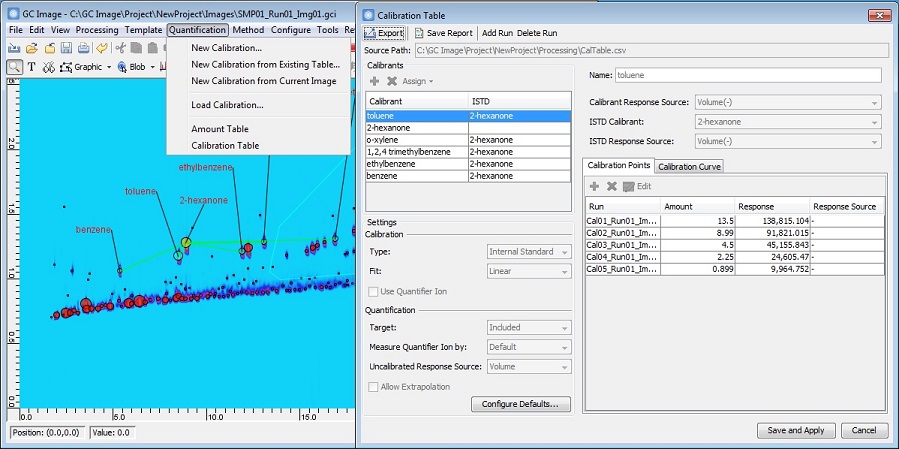
New Calibration and Quantification Tools
GC Image GCxGC Edition 2.7 Release 1 introduces new tools for calibration
and quantification. The process for generating calibration tables is
simplified by a new UI and options, including:
- New Quantification
menu options for tools to conveniently create calibration tables
directly from a set of chromatograms, and to load, edit, and save calibration tables.
- A new Calibration Table interface that allows users to examine
and edit individual calibrants. Calibration points and graphs of
the calibration curves are easily accessible through a tabbed view.
- New calibration function types, Quadratic and Piecewise Linear,
in addition to Linear.
- New calibration method files that allow calibration settings
to be customized and stored for reuse.
- Quantitation using ISTD Ions have been made easily available with the new
quantitation settings and UI improvements.
- Additional quantitation settings have been added to give users more
options. The quantification settings allow users to control the
targets to be quantified, how to measure the quantifier ions, the
uncalibrated response type, and if they want to allow extrapolation.
Contents Next: Introduction
GC Image™ GCxGC Edition User Guide © 2001–2017 by GC Image, LLC, and the
University of Nebraska.