GC IMAGE - GC Image GCxGC Edition - User Guide 2025R3
New Features and Improvements in Version 2.8
GC Image GCxGC Edition 2.8 has many new features and improvements. See
the GC Image GCxGC Edition User Guide for full
documentation.
What's New in Version 2.8 Release 3 (February 2019)
Version 2.8R3 has the following new improvements for
Processing > Extract Ion
Peaks as Area Objects:
- Check for existing ion peak areas before detection. The program will ask a
user to delete existing areas before detection. If the user chooses
not to delete them, the program will delete an existing ion peak area
automatically if an exact duplicate is detected.
- Supporting Gaussian or Savitzky-Golay smoothing for both
Column I and II;
- New Mass Range filter that allows a user to specify
one or more mass ranges (e.g. "41, 52.5-56.5");
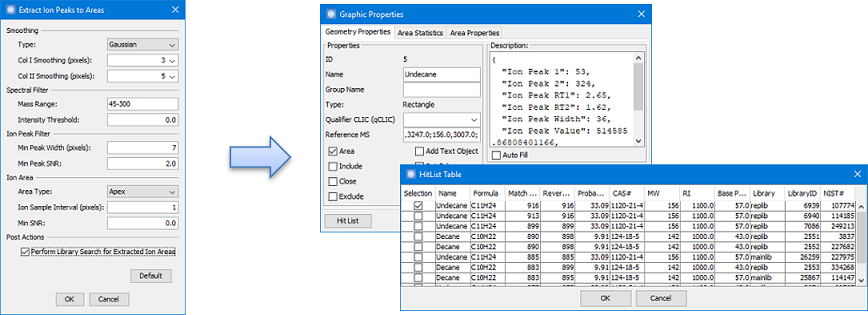
- New Perform Library Search for Extracted Ion Areas
option that turns on library search with the deconvolved spectra of
extracted ion peak areas automatically;
- Additional ion peak attributes stored as custom
attributes of the extracted ion peak areas including: Peak 1,
Peak 2, RT1, RT2, Peak Width, Peak Value,
and Qualifier Count. They can be added to Blob
Table through CLIC columns using CLIC function CUSTOMATTRIBUTE("Attribute
Name").
- The detection parameters are now recorded in Journal.
Version 2.8R3 has new improvements for Review mode including:
- All views are updated to display information of the currently
selected blob or area upon start review.
- During review, a blob or an area can be selected in Image View directly to
review its information. The selection action updates all views
including 1D View, SIC, Spectrum View, and Blob/Area Table selection
automatically.
Version 2.8R3 now includes replib from NIST Library in addition
to mainlib by default, in order to improve the accuracy of the
search results for common compounds.
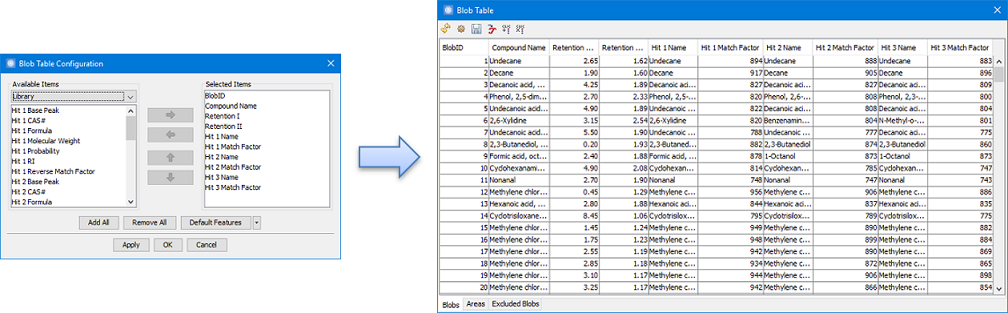
In addition, Blob
Table now has columns for accessing the top three hit list
entries returned by
Library Search including:
- Hit Name — Compound Name of the hit at the specified rank
- Hit Formula — Chemical Formula of the hit at the specified rank
- Hit Molecular Weight — Molecular Weight of the hit at the specified rank
- Hit Base Peak — Base Peak of the hit at the specified rank
- Hit CAS# — CAS Number of the hit at the specified rank
- Hit RI — RI of the hit at the specified rank
- Hit Match Factor — Match Factor of the hit at the specified rank
- Hit Reverse Match Factor — Reverse Match Factor of the hit at the specified rank
- Hit Probability — Probability of the hit at the specified rank
Additional hit list entries can be added to Blob Table as CLIC
columns, for example, "Hit_Name[5]" or "Hit_Match_Factor[5]".
Version 2.8R3 has new features for Investigator, including:
- %RSD (RSD in percents) added to computed
statistics in addition to absolute RSD.
- New Analysis menu that
replaces the configuration button for selecting computed statistics
and provides preset configurations of commonly used statistics
including Means, Overall Descriptive Statistics, Pairwise
Differences, and Multi-Class Statistics.
Version 2.8R3 has other new and improved operations, including:
What's New in Version 2.8 Release 2 (November 2018)
Version 2.8R2 now supports pulse calibration for
either centroid or profile data.
Custom attributes can now
be specified in the Description field of a blob or an area
using the JSON formatted name-value pairs. A new CUSTOMATTRIBUTE("Attribute
Name") CLIC
function can be used to retrieve custom attribute values to be used in
Blob Table for calculation and reporting.
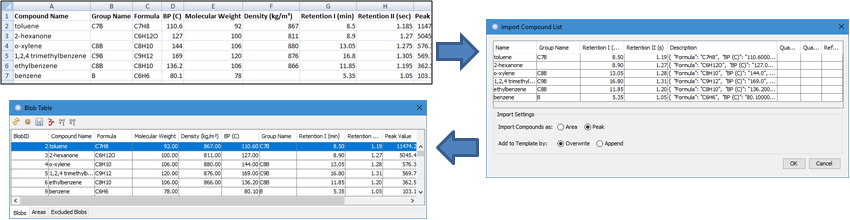
Template - Import
Compound List is enhanced to import both built-in and custom
attributes from external sources including:
- Support importing Group Name and Reference MS fields;
- Support importing Retention I and Retention II in either Minutes,
Seconds, or Pixels;
- Support importing multiple custom attributes to the
Description field.
Also, custom attributes for a sample can be added to a Sequence Table in a
project, and automatically imported into Image Attributes during auto
processing. A new MISCIMAGEATTRIBUTE("Attribute Name") CLIC
function can be used to retrieve custom attribute values from the Image
Attributes - Miscellaneous table to be used in Blob Table for
calculation and reporting.
Version 2.8R2 has new improvements for template
matching and raw data alignment including:
- Register Image
has a new option that allows re-detecting blobs that match the
pre-registration blobs automatically.
- Template -
Interactive Match and Transform now supports all transform types
including custom transform plug-ins.
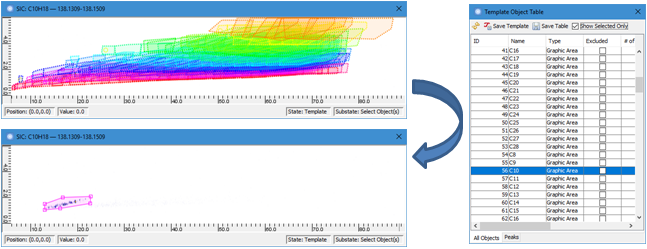
- Template
Object Table has a new Show Selected Only option. When the
option is selected, only selected template object(s) will be
displayed so that a user can more easily review and edit specific
template object(s), such as a compound group polygon in a complex
template.
GC Image program now has a new workspace mode if invoked
from GC Project program. When
Project launches an Image window instance with an open project, it
will set the Image window instance to use default paths of the project
for import, open, configuration, template, and report functions so
that a user can maintain all files related to a project together and
switch from one project to another easily.
Version 2.8R2 has other new and improved operations, including:
- The software now automatically detects the missing
MSFileReader dependency and notifies users when importing Thermo Raw
files.
- The Find Blob tool now stays open and
allows searching blobs continuously until it is closed by the user.
- Investigator now
shows the path of the open analysis file in its window title.
What's New in Version 2.8 Release 1 (July 2018)
GC Image GCxGC Edition 2.8r1 has new and improved I/O facilities,
including:
- The new MassHunter DAC B.08 that provides supports for the
new compression format for profile data, and IMS data.
- Improved support for the GCD files acquired by the new
Shimadzu LabSolution software (The support is experimental and limited
to GC data).
- The updated VUV plugin that supports data acquired by a
VGA 101 system.
- The software now automatically detects the missing Microsoft
Visual C++ runtime dependencies and notifies users for MassHunter and
Waters Raw data readers.
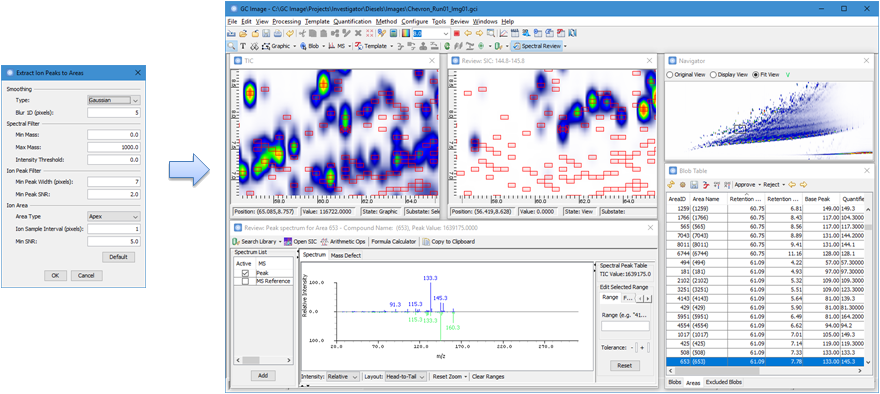
The new Processing >
Extract Ion Peaks as Area Objects replaces the Extract
Ion Peaks to Areas plugin included in the previous versions. The new
function includes several major improvements:
- A completely new algorithm that implements a processing
workflow similar to AMDIS with spectral deconvolution.
- A new implementation that is much faster and uses less
memory.
- Extracted ion peaks with their characteristic ions and
deconvolved spectra can be easily reviewed and identified through the
Review Spectrum view of the Review
Mode.
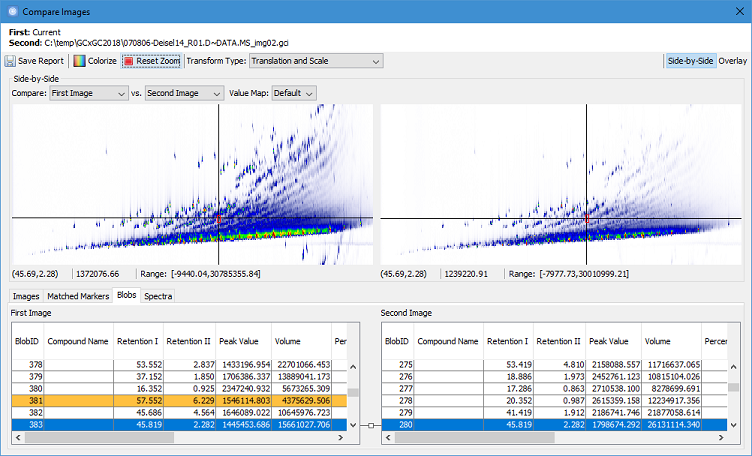
The new Compare Images -
Side-by-Side tool adapts the reliable peak region alignment methods
from Investigator, enables
comprehensive peak matching between two chromatograms, and automates
difference detection. The tool provides a set of visualizations that
allow users to interactively review two chromatograms, the alignment
transformation, and peak matches. It provides not only comparative
visualization, but also comprehensive characterization of sample
differences down to individual peaks.
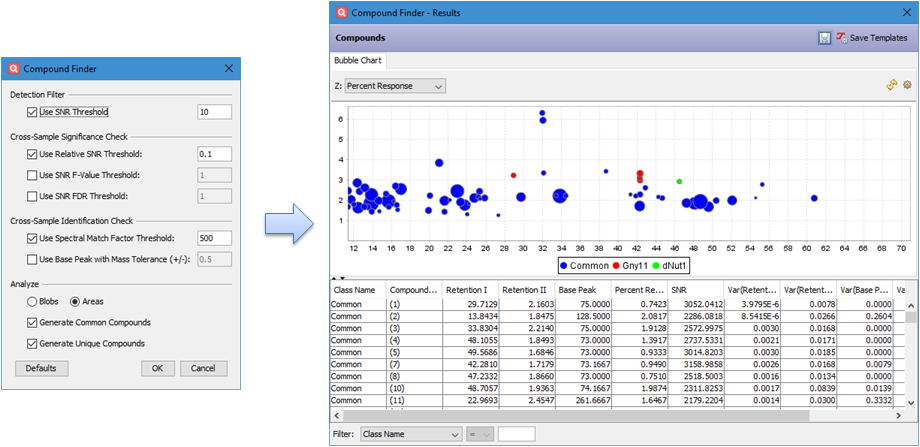
The new Compound Finder
tool in Investigator
allows analysts to detect common and unique compounds across many
samples with specialized detection and identification constraints that
use chromatographic and mass spectral information to distinguish
marker compounds. In addition, the results are visualized with a
labeled bubble plot, which provides not only metric values, but also
instructive predictions as to which features are effective for
distinguishing samples.
The new Investigator ML is a plugin tool for supervised pattern
recognition and classification. An experimental version of
Investigator ML is included with GC Image GCxGC Edition Version 2.8.
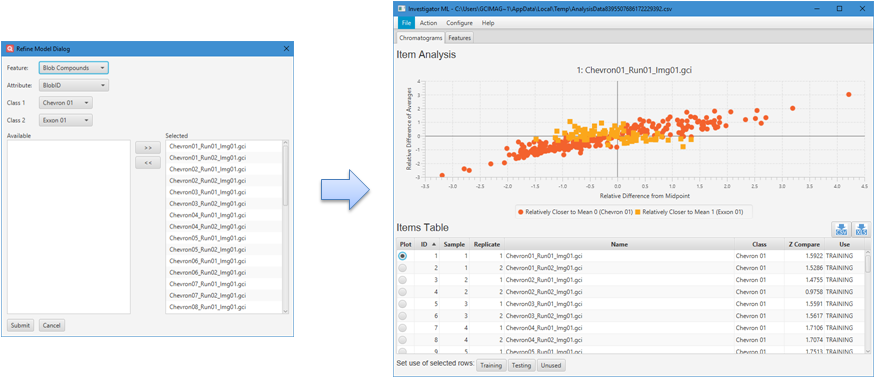
Investigator ML can be launched from the Investigator menu as Tools
> Investigator ML. Then, from the dialog, shown below, the user
selects the:
- Feature type, either blobs or areas;
- Attribute for pattern recognition, e.g., Percent Response;
- Classes for training and classification; and
- Chromatograms to provide feature data.
In the current version, Investigator ML supports only supervised
binary classification, so only two classes can be selected and only
chromatograms from those classes are available for selection as
sources of the feature data. Investigator ML has a separate Users'
Manual that describes its operations and use. This is available via
the Help menu in the Investigator ML interface.
Contents Next:
Introduction
GC Image™ GCxGC Edition User Guide ©
2001–2018 by GC Image, LLC, and the University of Nebraska.